
Contact information
Professor Robert Gilbert's Research Group
Henry Wellcome Building of Genomic Medicine
Colleges
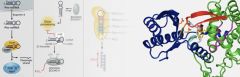 |
| miRNA maturation is switched by 3' uridylylation of pre-miRNA forms. We are studying this process using the yeast enzyme Cid1 and its close human homologues ZCCHC6 and ZCCHC11. The left hand region of the image shows how miRNAs are matured, including uridylylation. The right hand region shows the active site of Cid1 bound with UTP. ZCCHC6 and ZCCHC11 are important potential anti-cancer drug targets. |
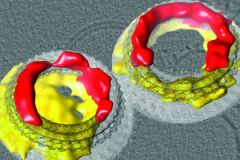 |
| Pores formed by membrane attack complex-perforin/cholesterol-dependent cytolysin proteins (MACPF/CDCs). These proteins are found in all different forms of cellular life and play key roles in infection and immunity. We recently showed that they can form pores constructed from proteins in arc-shaped oligomers and the membrane lipids themselves. This finding gives a critical insight into how these proteins act in the different diseases in which they are involved. |
Robert Gilbert
Professor of Biophysics
Our work is focused on molecular mechanisms underlying pathology in humans, specifically cancer and membrane pore formation and cell adhesion. We are studying mechanisms of 3' uridylation of RNAs with clear effects in tumourigenesis and are engaged in related translational research in collaboration with the Target Discovery Institute and Cancer Research Technology. We are also working on mechanisms of pore formation relevant to inflammation and cytotoxic T cell activity, and to infection processes in diseases such as malaria and toxoplasmosis. Our work on membrane biology in disease includes a focus on the kindlin proteins which are involved both in cell adhesion activation and also function nuclearly in suppression of tumour-suppressor miRNAs.
Data sharing statement: we deposit our results in relevant online databases where available for release on publication. Other outputs will be distributed on request.
Currently our work is being funded by Cancer Research UK, the British Heart Foundation, the Medical Research Council, the Biotechnology and Biological Sciences Research Council, and the Wellcome Trust.
Recent publications
The European Biophysics Journal Prizes 2025: Recognising biophysical science at sub-cellular, cellular and tissue levels of organisation.
Journal article
Gilbert RJC., (2026), European biophysics journal : EBJ, 55, 3 - 7
Covalently constrained ‘Di-Gembodies’ enable parallel structure solutions by cryo-EM
Journal article
Yi G. et al, (2026), Nature Chemical Biology, 22, 69 - 76
Structure and unusual binding mechanism of the hyaluronan receptor LYVE-1 mediating leucocyte entry to lymphatics
Journal article
Bano F. et al, (2025), Nature Communications, 16
Open architecture of archaea MCM and dsDNA complexes resolved using monodispersed streptavidin affinity CryoEM
Journal article
Ma J. et al, (2024), Nature Communications, 15
Structural basis for activity switching in polymerases determining the fate of let-7 pre-miRNAs
Journal article
Yi G. et al, (2024), Nature Structural & Molecular Biology, 31, 1426 - 1438


